Macvector editor1/13/2023 SPAdes in particular will optimize assembly based on these settings. Double-click on a read after importing into a project to set these parameters. There are now more options to control the type of read that you are using (unpaired, paired-end, mate pair, HQ mate pair, NxSeq long mate pair), or the source of the reads (Illumina, IonTorrent, PacBio, Nanopore etc). This dramatically reduces the amount of temporary disk space required when assembling large datasets. MacVector now directly supports compressed fasta and fastq formatted files (.gzip/.gz) and will submit those directly to assembly algorithms. MacVector will automatically identify them where possible. You can now use interleaved paired read files for all NGS assembly algorithms (velvet, bowtie and SPAdes). One drawback of SPAdes is that it does not generate alignments, only contig consensus sequences, so we have added an option to generate alignments within MacVector using bowtie. It is a little slower than our existing velvet algorithm, but does a better job of resolving repeats at the ends of contigs, so you generally find it produces longer contigs with the same input data. This is a high-performance assembler that uses very little RAM, allowing it to run on relatively modest machines, though we do recommend 16GB+ of RAM for optimal performance. We have added the popular SPAdes algorithm to Assembler for MacVector 16.0. 
If you want to change the appearance of multiple features at the same time, you will get the normal Symbol Editor. Note: this is only displayed when you are editing a single individual feature. This lets you edit both the basic Genbank-style feature information and the graphical symbol appearance in a single dialog. If you double-click on an individual feature in the Map tab, it now brings up a dialog with two tabs – one for the Feature Editor and one for the Symbol Editor. If you open a typical unannotated vector sequence, you will get a display something like this By default, MacVector uses the collection of features in the /MacVector/Common Vectors/Annotated Fragments/ folder, but you can use any folder you like, configured using the new MacVector | Preferences | Scan DNA preference settings. 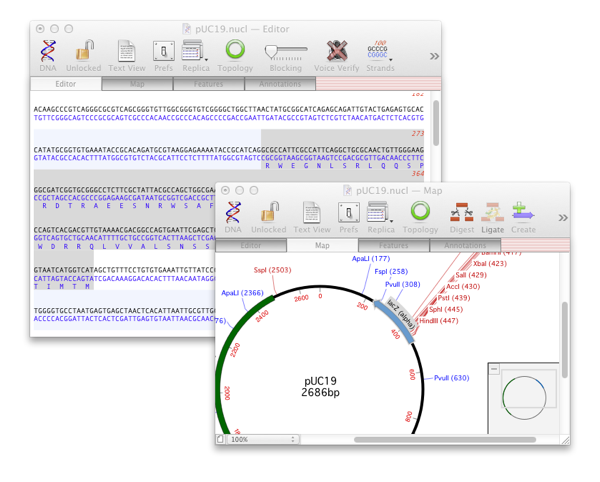
It shows these in faded colors in the Map tab so that you can see what features are missing from your sequence. 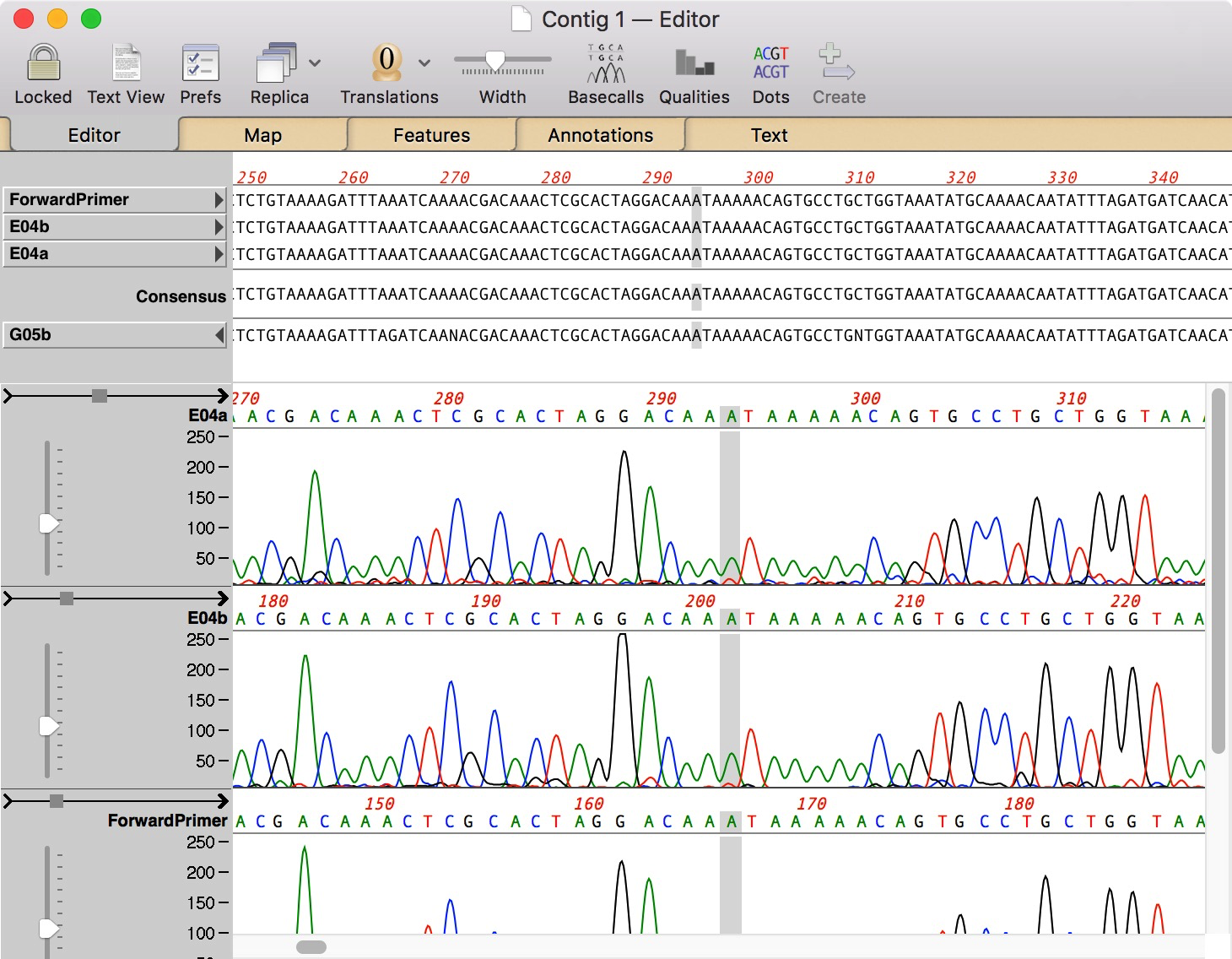
Whenever you open a DNA sequence, MacVector now scans it to identify common features that are not already annotated on the sequence. Assembler has a new de novo assembler algorithm ( SPAdes) as well as improvements to the handling and analysis of many different types of NGS data. There is also a new Feature Editor that lets you change the underlying "GenBank" data as well as the visual symbol appearance in the same dialog. This complements the existing searches for ORFs and Restriction Enzymes. MacVector 16 adds an automatic search for missing features whenever you open a DNA sequence. Sequence Analysis Tools for Molecular Biologists
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
